FOOOF - fitting oscillations & one over f¶
FOOOF is a fast, efficient, and physiologically-informed tool to parameterize neural power spectra.
Overview¶
The power spectrum model conceives of a model of the power spectrum as a combination of two distinct functional processes:
An aperiodic component, reflecting 1/f like characteristics, with
A variable number of periodic components (putative oscillations), as peaks rising above the aperiodic component
This model driven approach can be used to measure periodic and aperiodic properties of electrophysiological data, including EEG, MEG, ECoG and LFP data.
The benefit of fitting a model in order to measure putative oscillations, is that peaks in the power spectrum are characterized in terms of their specific center frequency, power and bandwidth without requiring predefining specific bands of interest and controlling for the aperiodic component. The model also returns a measure of this aperiodic components of the signal, allowing for measuring and comparison of 1/f-like components of the signal within and between subjects.
Documentation¶
Documentation is available on the documentation site.
This documentation includes:
Motivations: with motivating examples of why we recommend parameterizing neural power spectra
Tutorials: with a step-by-step guide through the model and how to apply it
Examples: demonstrating example analyses and use cases, and other functionality
API list: which lists and describes all the code and functionality available in the module
FAQ: answering frequency asked questions
Glossary: which defines all the key terms used in the module
Reference: with information for how to reference and report on using the module
Dependencies¶
FOOOF is written in Python, and requires Python >= 3.6 to run.
It has the following required dependencies:
There are also optional dependencies, which are not required for model fitting itself, but offer extra functionality:
matplotlib is needed to visualize data and model fits
tqdm is needed to print progress bars when fitting many models
pandas is needed to for exporting model fit results to dataframes
pytest is needed to run the test suite locally
We recommend using the Anaconda distribution to manage these requirements.
Installation¶
The current major release is the 1.X.X series, which is a breaking change from the prior 0.X.X series.
Check the changelog for notes on updating to the new version.
Stable Version
To install the latest stable release, use pip:
$ pip install fooof
The module can also be installed with conda, from the conda-forge channel:
$ conda install -c conda-forge fooof
Development Version
To get the current development version, first clone this repository:
$ git clone https://github.com/fooof-tools/fooof
To install this cloned copy, move into the directory you just cloned, and run:
$ pip install .
Editable Version
To install an editable version, download the development version as above, and run:
$ pip install -e .
Other Language Support¶
The original implementation of FOOOF, available in this repository, is implemented in Python.
If you wish to run FOOOF from another language, there are a couple potential options:
a wrapper, which allows for running the Python code from another language
a reimplementation, which reflects a new implementation of the fooof algorithm in another language
Below are listed some examples of wrappers and/or reimplementations in other languages (non-exhaustive).
Matlab¶
In Matlab, there is a reimplementation available in common toolboxes:
The Brainstorm toolbox has a reimplementation of fooof (see the Brainstorm fooof tutorial)
The Fieldtrip also uses the same reimplementation (see the Fieldtrip fooof tutorial)
There is also a Matlab wrapper in the fooof_mat repository.
Note that another option is to use Python FOOOF within a Matlab pipeline, as explored in the mat_py_mat repository.
Other Languages¶
Other languages with wrappers include:
Julia, for which there is a fooof wrapper
R, in which fooof can be run using reticulate, as shown here
Reference¶
If you use this code in your project, please cite:
Donoghue T, Haller M, Peterson EJ, Varma P, Sebastian P, Gao R, Noto T, Lara AH, Wallis JD,
Knight RT, Shestyuk A, & Voytek B (2020). Parameterizing neural power spectra into periodic
and aperiodic components. Nature Neuroscience, 23, 1655-1665.
DOI: 10.1038/s41593-020-00744-x
Direct link: https://doi.org/10.1038/s41593-020-00744-x
More information for how to cite this method can be found on the reference page.
Code and analyses from the paper are also available in the paper repository.
Contribute¶
This project welcomes and encourages contributions from the community!
To file bug reports and/or ask questions about this project, please use the Github issue tracker.
To see and get involved in discussions about the module, check out:
the issues board for topics relating to code updates, bugs, and fixes
the development page for discussion of potential major updates to the module
When interacting with this project, please use the contribution guidelines and follow the code of conduct.
Quickstart¶
This module is object oriented, and uses a similar approach as used in scikit-learn.
The algorithm works on frequency representations, that is power spectra in linear space.
Fitting a Single Power Spectrum
With a power spectrum loaded (with ‘freqs’ storing frequency values, and ‘spectrum’ storing the power spectrum, both as 1D arrays in linear space) FOOOF can be used as follows:
# Import the FOOOF object
from fooof import FOOOF
# Initialize FOOOF object
fm = FOOOF()
# Define frequency range across which to model the spectrum
freq_range = [3, 40]
# Model the power spectrum with FOOOF, and print out a report
fm.report(freqs, spectrum, freq_range)
FOOOF.report() fits the model, plots the original power spectrum with the associated FOOOF model fit, and prints out the parameters of the model fit for both the aperiodic component, and parameters for any identified peaks, reflecting periodic components.
Example output for the report of a FOOOF fit on an individual power spectrum:
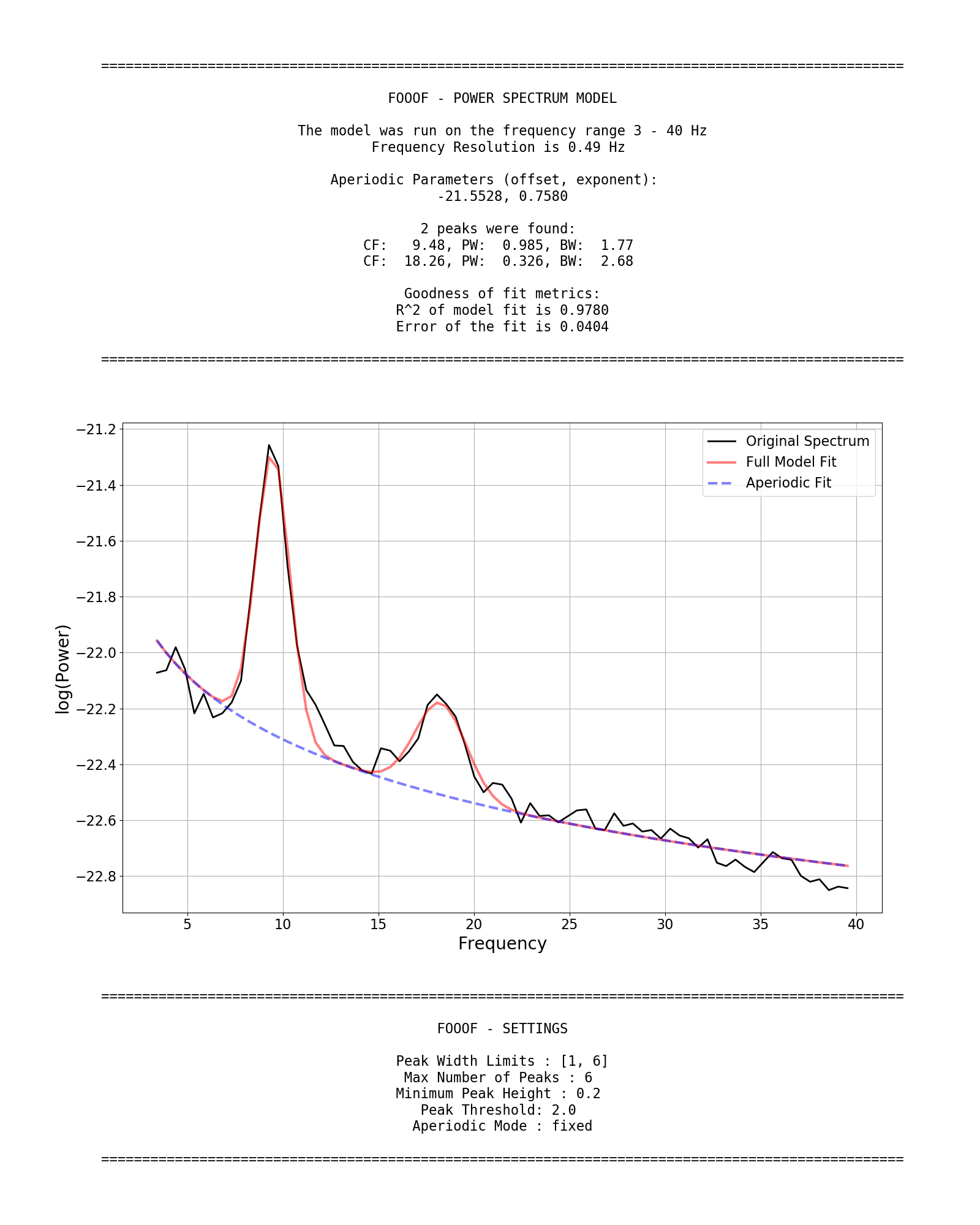
Defining the model Settings
The settings for the algorithm are:
peak_width_limitssets the possible lower- and upper-bounds for the fitted peak widths.max_n_peakssets the maximum number of peaks to fit.min_peak_heightsets an absolute limit on the minimum height (above aperiodic) for any extracted peak.peak_thresholdsets a relative threshold above which a peak height must cross to be included in the model.aperiodic_modedefines the approach to use to parameterize the aperiodic component.
These settings can be defined when initializing the model, for example:
# Initialize a FOOOF model object with defined settings
fm = FOOOF(peak_width_limits=[1.0, 8.0], max_n_peaks=6, min_peak_height=0.1,
peak_threshold=2.0, aperiodic_mode='fixed')
Fitting a Group of Power Spectra
Next is an example workflow for fitting a group of neural power spectra. In this case, ‘freqs’ is again a 1D array of frequency values, and ‘spectra’ is a 2D array of power spectra. We can fit the group of power spectra by doing:
# Initialize a FOOOFGroup object, specifying some parameters
fg = FOOOFGroup(peak_width_limits=[1.0, 8.0], max_n_peaks=8)
# Fit FOOOF model across the matrix of power spectra
fg.fit(freqs, spectra)
# Create and save out a report summarizing the results across the group of power spectra
fg.save_report()
# Save out FOOOF results for further analysis later
fg.save(file_name='fooof_group_results', save_results=True)
Example output from using FOOOFGroup across a group of power spectra:
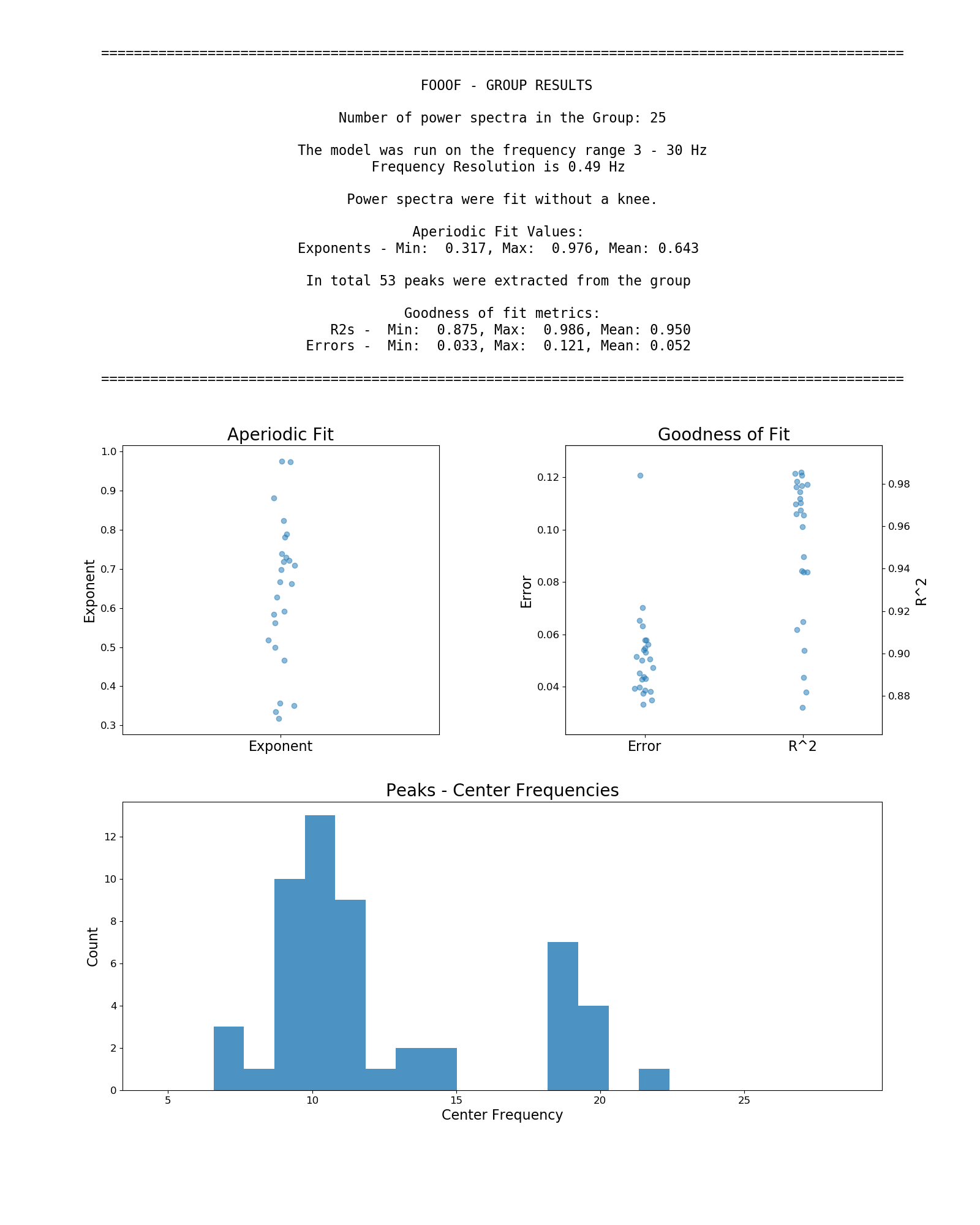
Other Functionality
The module also includes functionality for fitting the model to matrices of multiple power spectra, saving and loading results, creating reports describing model fits, analyzing model outputs, plotting models and parameters, and simulating power spectra, all of which is described in the documentation.
Funding¶
Supported by NIH award R01 GM134363 from the NIGMS.




